Our research
We study how cells make a perfect copy of the genome, a process called DNA replication.
Failures in this process
are the root cause of many diseases, particularly cancers, and inhibitors of DNA replication are
frequently used in the
clinic as anti-viral agents and chemotherapies. Our studies shed light on the consequences of these drugs
for cell division control.
DNA replication control during the cell cycle
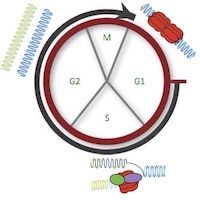
It is vitally important that the entire genome is copied perfectly in every cell division. Failures in this copying process – termed DNA replication – result in mutations that are the root cause of cancers. In our lab we study how perfect DNA replication is achieved during a single cell cycle. We work primarily with the budding yeast Saccharomyces cerevisiae and we apply our findings to other systems including cancer cells.
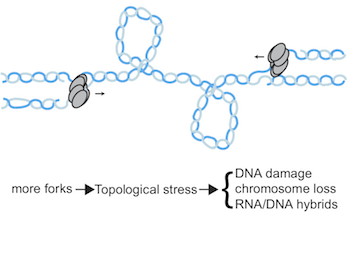
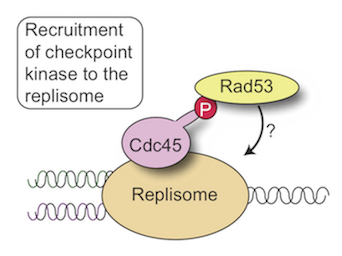
DNA damage and genome stability
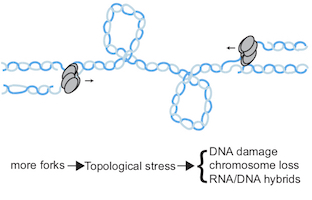
Difficulties during genome duplication, such as a lack of metabolites, are a common feature of cancer and ageing cells. Such replication stress activates a checkpoint kinase response, which is essential for rescuing genome duplication and are an active area of interest for the development of new chemotherapeutic agents. We are interested in understanding the mechanisms and functions of this checkpoint kinase response.
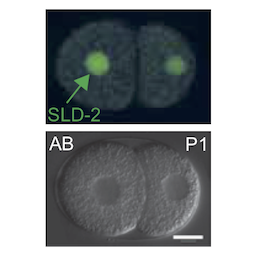
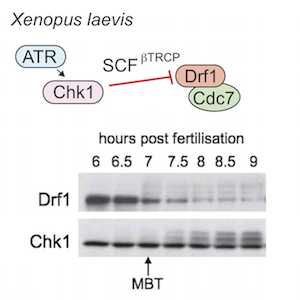
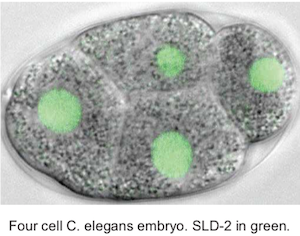
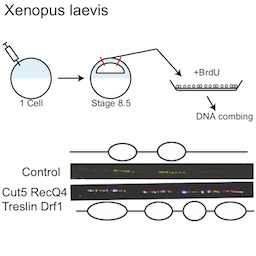
Genome duplication and development
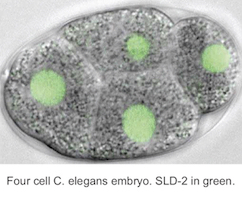
Healthy tissues coordinate cell proliferation with differentiation and for many cell types this involves dramatic changes in genome duplication. The mechanisms and physiological importance of these changes in DNA replication are very poorly understood. We study C. elegans and Xenopus embryos to understand the function of S-phase control during healthy animal development.
Public engagement and outreach

As part of the University of Cambridge and Wellcome Trust commitment to public engagement and widening participation, we are actively involved in school visits and public interactions with our research. Here is a picture of Philip Zegerman before giving a talk at the Antony Gormley exhibition at Kettle’s yard.